HYDROID
HYDROID (HYDroxyl-Radical fOotprinting Interpretation for DNA) is a python package for the analysis of the experimental data generated by hydroxyl-radical footprinting (HRF) of DNA-protein complexes and its interpretation through comparison to theoretical predictions from molecular models.
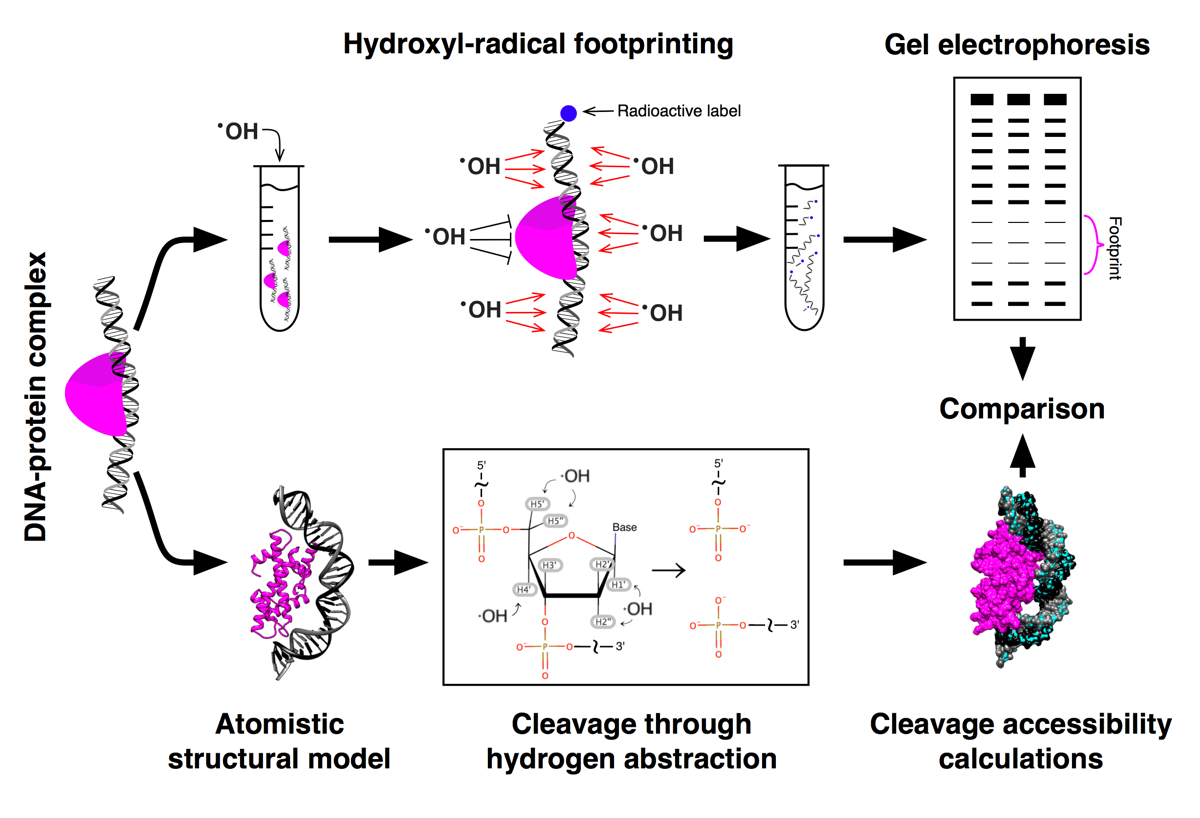
HYDROID features
- 2 in 1: HRF experimental data quantification + theoretical analysis of atomistic structures
- Extracts cleavage intensities at individual DNA nucleotides by a number of constraint fitting algorithms
- Uses both Gaussian and Lorentzian models for band intensities
- Cross-platform python-scripted solution, can be install on Linux, MacOS, Windows
- Completely free and relies on open source components such as ImageJ and FreeSASA
- Provides examples of raw data analysis together with data analysis workflows.
Documentation
For detailed documentation - click here.
HYDROID video tutorial is available here.
Quick-start guide
NOTE: HYDROID is a full featured Python-script driven software solution that requires basic familiarity with Python-scripting.
HYROID_GUI is a sister package that wraps some basic gel lane quantification functionality into a more user friendly graphical interface. HYDROID_GUI video tutorial is available here.
Quick installation
Install Miniconda with Python2.7 for your platform from https://conda.io/miniconda.html.
NB: As of 2020 only the legacy version 4.3.31 of the Miniconda installer works that can be downloaded from https://repo.anaconda.com/miniconda/
conda install -c hydroid hydroid
Test HYDROID:
HYDROID_test_exp #Tests exeprimental data analysis module
HYDROID_test_pred #Tests molecular structure analysis module (currently supported on Linux and OSX)
For alternative installation instructions for Linux, MacOS and PC see INSTALL.md.
Start by downloading and modifying an example
HYDROID_get_ex1
cd example1
python exp_s2_assign_peaks.py
...
See full examples set and instructions in examples folder.
Citing HYDROID
Please cite HYDROID using following publication:
- A.K. Shaytan, H. Xiao, G.A. Armeev, D.A. Gaykalova, G.A. Komarova, C. Wu, V.M. Studitsky, D. Landsman, A.R. Panchenko “Structural interpretation of DNA–protein hydroxyl-radical footprinting experiments with high resolution using HYDROID”, Nature Protocols, 2018, DOI: 10.1038/s41596-018-0048-z